WP 1: Vibrio – ecosystem engineer relations in the past
The distribution of eelgrass (Zostera marina) changed during the last decades in the Baltic Sea. Large-scale declines of eelgrass have been reported from areas of low water clarity and high nutrient concentrations. Similarly, also macrophytes and blue mussel beds have undergone changes in coverage, recruitment and/or depth distribution.
Recordings of Vibrio infections in the Baltic Sea region are handled differently by the Baltic countries, and neither coordinated Vibrio monitoring nor harmonized databases exist that could provide an overview of Vibrio cases. In principle data sets exist, some of which date back several decades, but up to now Vibrio infections have never been associated with mussel beds, macroalgae, or seagrass meadow occurrences. The aim of WP1 is to generate an overview of all known Vibrio cases of the past for the entire Baltic Sea region and to relate them to the presence of these ecosystem engineers at the time of infection. The data will be used to test and evaluate the results of WPs 2 and 3.
Main aims of WP 1:
• Identification and mapping of pathogenic Vibrio presence in the Baltic Sea.
• Linking cases of Vibrio infection with water temperature and salinity.
• Linking Vibrio infection and the presence of mussel beds, macroalgae, or seagrass.

WP 2: Vibrio – current distribution, regulation and pathogenicity
The objective of WP2 is to determine the current in situ pelagic and benthic (and biofilm) Vibrio composition under different environmental conditions and in relation to characteristic coastal habitats such as eelgrass meadows, macroalgae or blue mussels beds. Research cruise-based as well as local sampling will be carried out from a large number of coastal sites along a salinity gradient from 25 (coast of Denmark) to <3-6 (Finland and Estonia). Additionally, time-series sampling will be carried out at selected sites. Vibrio analyses will be conducted both by cultivation and by molecular analyses. Approx. 100 Vibrio isolates will be whole-genome sequenced to gain further insight into the genetic diversity and pathogenic potential of Baltic Sea Vibrios. Based on publicly available genomes and the new genomes sequenced in this study, Vibrio-specific pathogenicity genes will be identified to design quantitative PCR (qPCR) assays for in situ quantification (for use here and in WPs 3 and 4). The “Vibrio-relative abundance index” (VAI) will be used to describe the Vibrio-bacterioplankton ratio by qPCR. In situ pathogenic Vibrio abundances will be determined using known pathogenicity genes and based on the results of the Vibrio isolate sequencing. The molecular Vibrio analyses will be accompanied by general bacterial community composition analyses to investigate potential bacteria-bacteria interactions. Overall, the data, in addition to already available data (WP1), will give insights into the current distribution of Baltic Vibrio spp. in the presence and absence of the three ‘ecosystem engineers’ and in relation to environmental conditions (e.g. temperature, salinity, nutrient and carbon levels).
Main aims of WP 2:
• In situ sampling of Vibrio.
• Cultivation and sequencing of Vibrio.
• Determining in situ distribution and regulation of Vibrio species.

WP 3: Projected Vibrio - ecosystem engineer relations
WP3 is primarily experimental and will manipulate parameters identified as main drivers of in situ Vibrio prevalence, e.g. temperature, salinity, water column nutrients, allochthonous carbon, the presence and absence of ecosystem engineers. In particular, we will investigate how climate change will affect the abundance of pathogenic Vibrio spp. in representative Baltic Sea areas. Basis for the experiments are the summer in situ assemblages of total bacterioplankton and conditions as determined in WP2. Micro- and mesocosm experiments testing effects of future (2050-2100) climate scenarios will be performed. Simplified benthic hard- and soft-bottom community assemblages will be tested for their effects on bacterioplankton with or without blue mussels (Mytilus spp.), macrophytes (Fucus vesiculosus) and eelgrass (Zostera marina). For relevant bacteria strains isolated during the experiments, pathogenic and non-pathogenic subgroups will be identified. The assumed impact of climate change scenarios on ecosystem engineers/Vibrio relations will be tested and evaluated using results of WPs 1 and 2.
Main aims of WP 3:
• Microcosm experiments to identify environmental driver combinations.
• Outdoor mesocosm experiments testing the effects of future Baltic Sea scenarios on pelagic and benthic Vibrio prevalence.
• Synthesis of climate change effects on ecosystem engineers/Vibrio relations.
WP 4: Potential of underwater islands as an NbS to reduce pathogenic Vibrio spp.
Preliminary studies have shown that ecosystem engineers (e.g. eelgrass, macrophytes, mussel beds) can keep Vibrio abundances in the surrounding water at a low level. The objective of WP4 is to test mobile underwater carriers, which can be anchored to the sediment. These can principally be overgrown by ecosystem engineers and be used flexibly in coastal environments. WP4 will evaluate the impacts of those overgrown islands in vitro and in situ in order to assess the applicability of these ‘mobile islands’ for marine coastal ecosystems (together with WP5). Vibrio sampling and analyses will be performed as described in WP2.
Main aims of WP 4:
• Determining the impact of commercial underwater islands on coastal microbial populations.
• Determining influence of naturally overgrown underwater islands on Vibrio in vitro.
• Determining influence of naturally overgrown islands on Vibrio in situ.
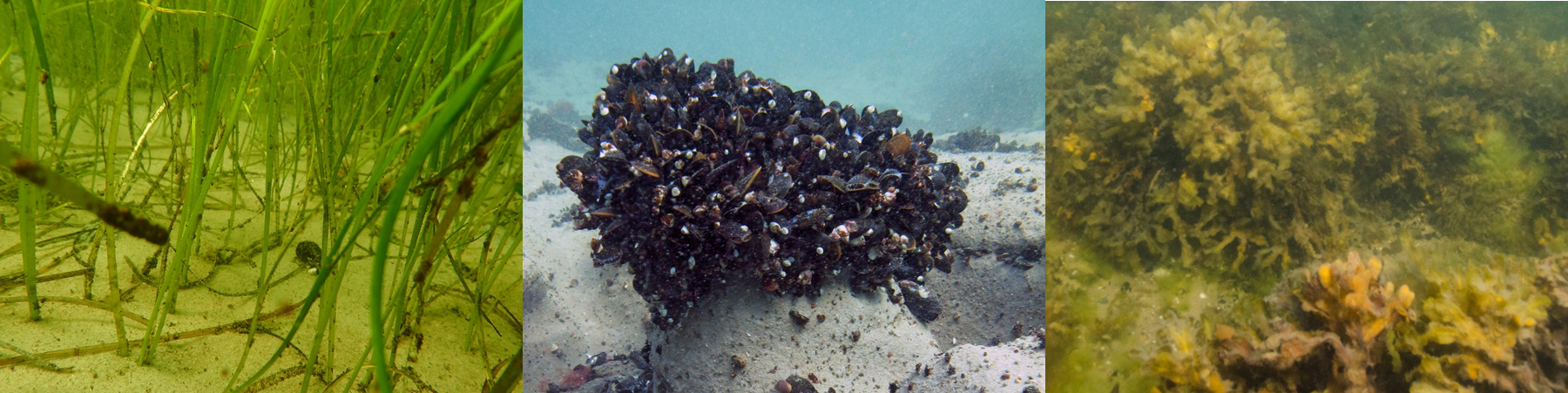

WP 5: Socio-ecological strategies
Long-term economic sustainability of coastal communities is dependent upon coastal ecosystems and the services they provide; i.e. if a health risk is associated with swimming in the future, due to Vibrio, it will be detrimental for the tourism in large coastal regions. BaltVib will include a network of Baltic researchers and stakeholders right from the project start and equip regulatory bodies and coastal authorities with the knowhow to accommodate the negative economic and environmental implications caused by the future Vibrio health risk. WP5 will integrate results from WPs 1-4 and produce geographical maps describing the ecosystem engineers-related composition, abundance, and infection potential of Vibrio spp. for the years 2021/2022 and 2100 for the investigated Baltic Sea areas. Institutions responsible for coastal monitoring and management will receive scientific support to prepare strategies to mitigate the Vibrio problem; e.g., by informing stakeholders how identified nature based solutions could reduce the risks.
Main aims of WP 5:
• Identification and mapping of societal and political stakeholders.
• Evaluation of stakeholder’s need regarding knowledge of the problem.
• Assessment of the impact of biodiversity changes on human health and economic valorization of nature based solutions.
• Knowledge exchange and capacity building with regional stakeholders.
• Dissemination of applied products to stakeholders and policy makers.
WP 6: Project and data management
The project coordinator will 1) monitor timely accomplishment of tasks and deliverables, and 2) ensure dissemination of the project outcome to relevant institutions and policymakers (together with WP5). Active and well-structured management will be based on close communication between the coordinator, the management board (all PIs), and a project advisory group composed of relevant scientists and stakeholders from all partner countries. This will happen through annual project meetings and regular video conferences to evaluate progress and address pertinent questions. The IOW will coordinate the samplings and experiments, including exchange of results and analyses, expertise, methodology and personnel between partners. As part of project management, a homepage will be developed and maintained.
Main aims of WP 6:
• Project management.
• Meetings and communication.
• Data management.