Aquatic monitoring using microbial nucleic acids
The challenge: One of the most pressing human problems is the loss of biodiversity and ecosystem functions in intensively used ecosystems. Anthropogenic factors play a dramatic and central role in destabilizing ecosystems and range from the input of nutrients and pollutants into rivers and lakes via agricultural use and wastewater to factors such as climate-active emissions and the extensive use of landscape. However, the scientific study of how potential environmental pollutants might affect entire ecosystems is not straightforward. For example, the highly dynamic, complex nature of ecosystems severely limits the effectiveness of experimentation and mathematical modelling; we are often left only with the means of environmental monitoring. In addition, the measurement of specific pollutants is often so laborious and costly that it is rarely possible in practice to observe a broad class of chemical pollutants in longer-running environmental monitoring programs.
The approach: The interconnectedness of organisms in an ecosystem is both a blessing and a curse: While it complicates many research approaches, it creates the opportunity to use individual organisms or groups of organisms like measuring devices for overall ecosystem health as well as other ecosystem parameters. As part of OTC Genomics, we will investigate whether and how we can use the microbiome as an indicator of the pollution of the Warnow River and the Baltic Sea coast in the Rostock-Heiligendamm area by various herbicides, pharmaceuticals and UV filters. We capture the microbiome's composition by high-throughput sequencing of the 16s and 18s rRNA genes; to analyse the resulting very large amounts of data, we employ cutting-edge machine learning and deep learning methods.
The great advantage: Whereas previous methods of environmental monitoring often focused on specific, morphologically well-distinguishable microorganisms such as diatoms and dinoflagellates, sequence-based biomonitoring enables a more taxonomically diverse view into ecosystems -- and thus likely much more accurate estimates of the anthropogenic pressures the ecosystems are facing. One of the goals of this research project is to achieve a high degree of automation so that almost no manual intervention is required -- from autonomous sampling to laboratory processing of samples to visual display of the collected data.
An automatic flow injection sampler
The challenge: Microbial communities are the main drivers of biogeochemical cycling of multiple elements sustaining life in the ocean. The rapidity of their response to stressors and abrupt environmental changes implies that even fast and infrequent events can affect local transformations of organic matter and nutrients. One of the promising approaches to investgate the function of microbial communities is the analysis of the transcripts in natural microbial assemblages (metatranscriptomes). Unfortunately, transcripts can degrade in less than 30 sec. Their unbiased detection in nature, especially from hypoxic or deep water habitats, is a challenge because they are subject to considerable modification simply due to sampling procedures. Modern molecular techniques now allow for monitoring of microbial activities and functions in the environment through the analysis of structure and function of natural microbial assemblages (Charvet et al., 2019).

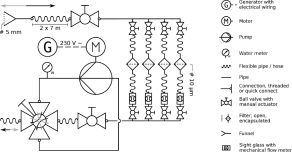
The solution: The first generation of a new Automatic Flow Injection Sampler (AFIS) was developed in our working group in 2012 (Feike et al. 2012). This AFIS samples and fixes water directly in the environment, and by this instantaneously conserves the gene expression profile in-situ. However, for a high resolution and standardized monitoring of temporal variations autonomous in situ fixation instruments are essential. This next generation AFISsys was developed in the frame of the BONUSproject AFISmon. AFISsys is a time - programmable, autonomous multi water sampler. Each prototype of AFISsys features several independent removable sample chambers that can collect up to 200 ml of water each. Every sampling chamber is automatically filled with a selected volume of fixative fluid solution after the water sample has been autonomously collected by the instrument itself.

The great advantage: AFISsys is capable of immediate fixation of in-situ collected water samples that can be conserved for up to one week before being processed for later analysis in the laboratory. This capability renders it an unique instrument for solving the issue of the inevitable delay between the collection of the sample and the extraction of its nucleic acids (DNA and RNA), allowing the investigation of the structure and the functions of microbial communities (meta-omics analyses) by limiting the bias effects of sample processing procedures.



Development of an encapsulated filtration device for microplastic sampling

The challenge: Microplastics are ubiquitous in the environment, including aquatic ecosystems. However, the qualitative and quantitative determination of microplastics in aquatic systems is methodologically complex. On the one hand, the small size of the particles poses the risk of sample loss, and on the other hand, contamination of the sample with plastics by the sampling equipment and the type of sample processing must be avoided.
The solution: In the working group Environmental Microbiology, we have developed a mobile sampling device for field sampling as part of the BMBF-funded project "Microcatch_Balt", which meets the requirements for obtaining contamination-free quantitative microplastic samples with a lower size limit of 10 µm using an encapsulated flow-through filtration concept (Lenz et al. 2018).
The great advantage: This new system "Rocket" offers two main advantages. With the filters connected in parallel in the closed system, all particles larger than 10 µm are collected. In addition, with the exception of the relatively rare PTFE (polytetrafluoroethylene), no plastic has been used in the device. This means that sampling can be assumed to be virtually contamination-free.