
Dr. Hagen Radtke - Research Interests
Decadal variability and long-term trends in Baltic Sea salinity
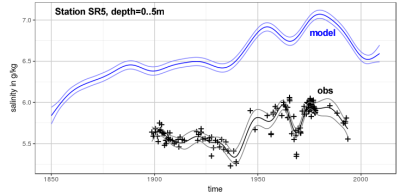
Climate projections suggest a decrease in Baltic Sea salinity. But in how far are the drivers influencing the Baltic Sea salinity represented correctly in Baltic Sea models? To answer this question, we perform historical recontructions (1850-present) with our hydrodynamical models to answer the question how much of the observed decadal variability in the salinity is reproduced by the models. The model results then allow a detailed investigation and and attribution of the drivers of these long-term changes.
Mechanistic modelling of sediment processes in marine ecosystems
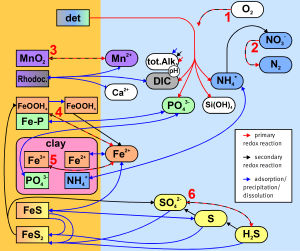
Biogeochemical processes in the sediments play a major role for the ecosystem and its nutrient balance, especially in shallow marine areas. In present-day ecosystem models, these processes are, however, highly parameterised, i.e. important quantities such as the ratio between remineralisation and burial of sedimented organic material are prescribed to the model based on measurements of the present-day state. This is obviously a drawback when the models are applied for future projections. We therefore strive to complement our ecosystem model ERGOM by a mechanistic sediment component (early diagenetic modelling).
Element tagging as a diagnostic tool in ecosystem models
In reality, atoms of the same element are mostly indistinguishable. An exception are istopic studies which play a major role in empirical investigations of ecosystem processes. In the model world, such a possibility to distinguish atoms can be added artificially, such that e.g. phosphorus can be "tagged" to identify its source (e.g. a specific river). But also elements which were involved in specific ecosystem processes can be tagged, in this way generating new possibilities to gain insights to what happens in the ecosystem model. Past applications of this method include the investigation of transport pathways of nitrogen and phosphorus from different rivers, quantifying effective nitrogen exchange between North Sea and Baltic Sea and investigating the question how important entrainment of ambient oxygen into deepwater inflows is for the oxygenation of the Baltic deep basins.
Numerics of differential equations in ecosystem models
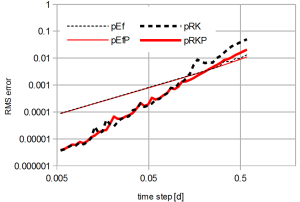
Biogeochemical processes in an ecosystem model are typically described by a system of ordinary differential equations. Numerical solvers for these systems should have some desirable properties. We developed the first numerical scheme which satisfies the following reqirements simultaneously:
- Multi-element conservation (the mass of several elements (e.g. nitrogen and phosphorus) needs to be conserved exactly)
- Positivity (no negative concentrations)
- Second-order accuracy
- Explicit time stepping scheme (no need to perform expensive matrix inversions)
- Ability to solve selected stiff ODE systems
Code Generation Tool for ecosystem models
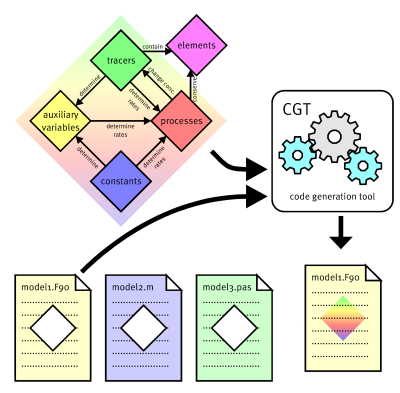
Ecosytem model source code is typically a mix of the scientific core and technical aspects. The scientific core are differential equations describing how the values of a state variable (e.g. the biomass of a functional group) change depending on environmental conditions. The technical environment e.g. requires allocation of arrays or forwarding results to an output management routine. The Code Generation Tool (www.ergom.net) which I developed makes it possible to separate the scientific core from the technical data handling. The source code is then automatically generated based on a formal description of the model in a set of text files. This has a lot of advantages, e.g.:
- Using the same ecosystem model in different physical host models (e.g. 1-d MATLAB test model, 3-d Fortran production model)
- Automatic check of element conservation
- Automatic element tagging
- Development of ecosystem models by non-programming experts possible
The Code Generation Tool is Open Source Software and is also applied outside IOW.
Efficient model validation
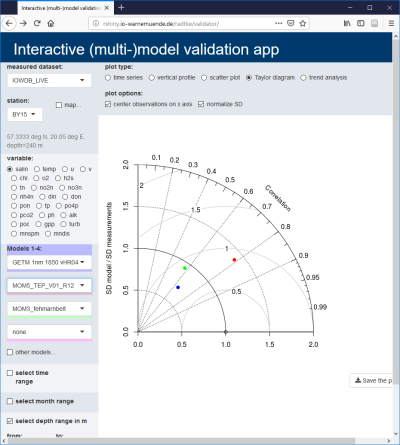
A thorough model validation is an essential prerequisite for the model's applicability to research questions. A standard evaluation method is often insufficient, since different variables or spatial areas need to be considered depending on the research question. The R Shiny app "Validator" allows a web-based validation of models against measurements which are live-extracted from the IOW database. Different plot types (time series, depth profile, Taylor diagram) can be created with a few clicks within seconds. A special advantage is the fact that model output can be checked during runtime already, so model runs deviating too far from the observations can be cancelled, saving valuable computation time.